Soybean Seed Tissues/Compartments GeneChip Experiments
We completed 91 GeneChip experiments of 40 tissues/compartments throughout soybean seed development using Affymetrix soybean IVT array (click here to view our annotation of soybean IVT array).
Figure. Tissues/Compartments Studied Throughout Soybean Seed Development (All images were hand drawn by Sharon Lee Belkin(Morphographics). The drawings of soybean seed are not to scale. Click on the image to enlarge.)
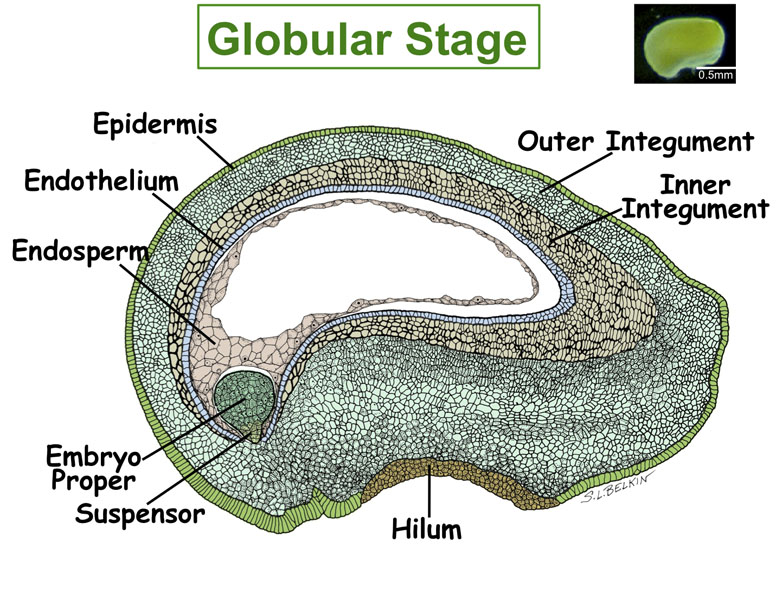 |
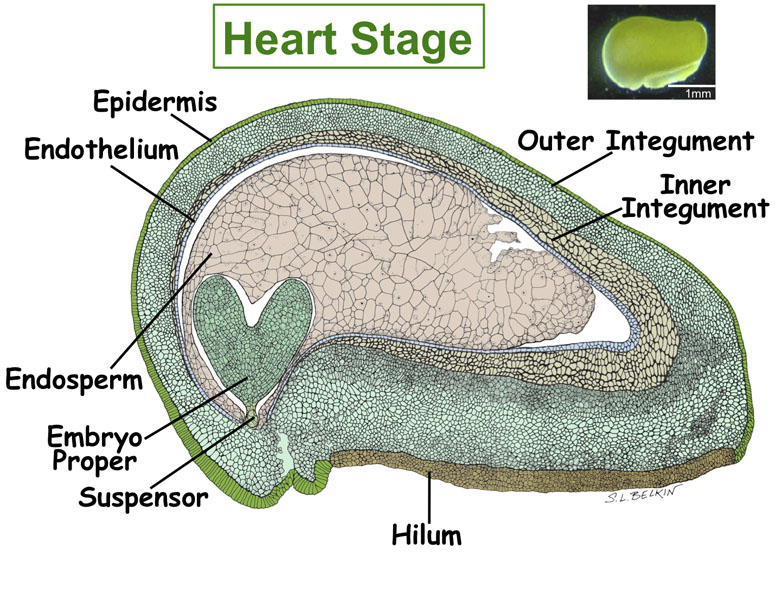 |
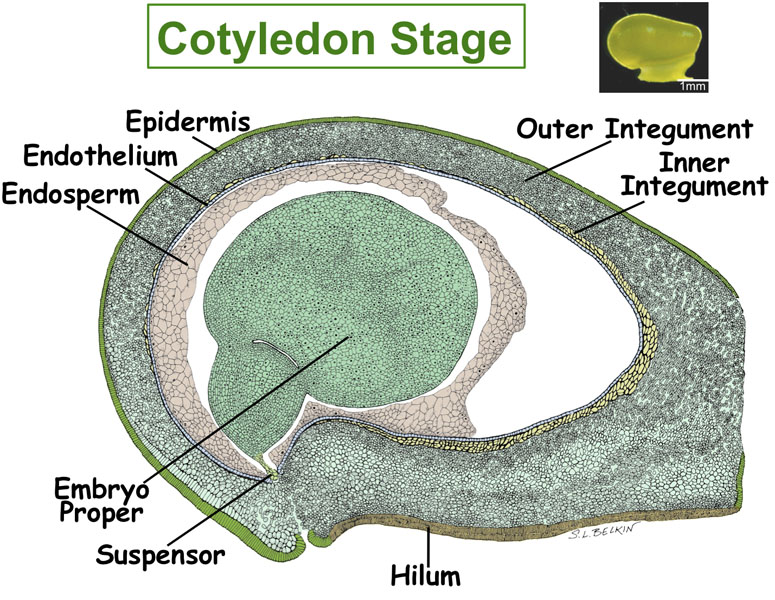 |
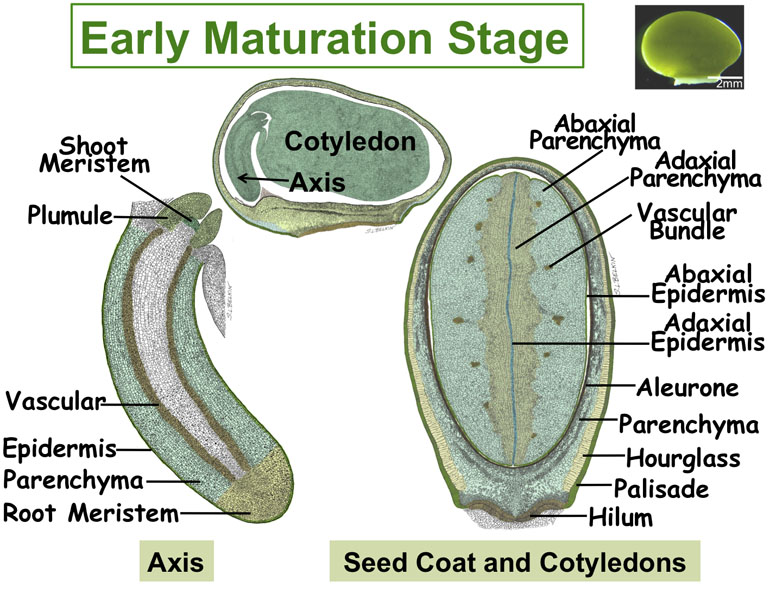 |
Click here to use our web-based analysis tools to analyze GeneChip experiments of soybean seed tissues/compartments.
Click on the name of soybean seed tissues/compartment listed below to find the detail information about GeneChip experiments of soybean seed tissues/compartments. The information includes:
(1) Soybean growth conditions;
(2) Protocols of RNA isolation and amplification;
(3) Images of section captured;
(4) GeneChip hybridization & analysis settings;
(5) Gene Expression Omnibus(GEO) accession number.
The notation for the experiment is as follows: GeneChip Array / Plant Species / Developmental Stage / Cell & Tissue Type.
- Soybean Array/Soybean/Cotyledon Stage/Embryo Proper
- Soybean Array/Soybean/Cotyledon Stage/Endosperm
- Soybean Array/Soybean/Cotyledon Stage/Endothelium
- Soybean Array/Soybean/Cotyledon Stage/Epidermis
- Soybean Array/Soybean/Cotyledon Stage/Hilum
- Soybean Array/Soybean/Cotyledon Stage/Inner Integument
- Soybean Array/Soybean/Cotyledon Stage/Outer Integument
- Soybean Array/Soybean/Cotyledon Stage/Seed
- Soybean Array/Soybean/Cotyledon Stage/Suspensor
- Soybean Array/Soybean/Early Maturation Stage/Aleurone
- Soybean Array/Soybean/Early Maturation Stage/Axis-Epidermis
- Soybean Array/Soybean/Early Maturation Stage/Axis-Parenchyma
- Soybean Array/Soybean/Early Maturation Stage/Axis-Plumule
- Soybean Array/Soybean/Early Maturation Stage/Axis-Root Meristem
- Soybean Array/Soybean/Early Maturation Stage/Axis-Shoot Meristem
- Soybean Array/Soybean/Early Maturation Stage/Axis-Vascular Bundle
- Soybean Array/Soybean/Early Maturation Stage/Cotyledon-Abaxial Epidermis
- Soybean Array/Soybean/Early Maturation Stage/Cotyledon-Abaxial Parenchyma
- Soybean Array/Soybean/Early Maturation Stage/Cotyledon-Adaxial Epidermis
- Soybean Array/Soybean/Early Maturation Stage/Cotyledon-Adaxial Parenchyma
- Soybean Array/Soybean/Early Maturation Stage/Cotyledon-Vascular Bundle
- Soybean Array/Soybean/Early Maturation Stage/Seed
- Soybean Array/Soybean/Early Maturation Stage/Seed Coat-Hilum
- Soybean Array/Soybean/Early Maturation Stage/Seed Coat-Hourglass
- Soybean Array/Soybean/Early Maturation Stage/Seed Coat-Palisade
- Soybean Array/Soybean/Early Maturation Stage/Seed Coat-Parenchyma
- Soybean Array/Soybean/Globular Stage/Embryo Proper
- Soybean Array/Soybean/Globular Stage/Endosperm
- Soybean Array/Soybean/Globular Stage/Endothelium
- Soybean Array/Soybean/Globular Stage/Epidermis
- Soybean Array/Soybean/Globular Stage/Hilum
- Soybean Array/Soybean/Globular Stage/Inner Integument
- Soybean Array/Soybean/Globular Stage/Outer Integument
- Soybean Array/Soybean/Globular Stage/Seed
- Soybean Array/Soybean/Globular Stage/Suspensor
- Soybean Array/Soybean/Heart Stage/Embryo Proper
- Soybean Array/Soybean/Heart Stage/Endosperm
- Soybean Array/Soybean/Heart Stage/Endothelium
- Soybean Array/Soybean/Heart Stage/Epidermis
- Soybean Array/Soybean/Heart Stage/Hilum
- Soybean Array/Soybean/Heart Stage/Inner Integument
- Soybean Array/Soybean/Heart Stage/Outer Integument
- Soybean Array/Soybean/Heart Stage/Seed
- Soybean Array/Soybean/Heart Stage/Suspensor
- Soybean Array/Soybean/Ovule Stage/Embryo Sac
- Soybean Array/Soybean/Ovule Stage/Integuments
- Soybean Array/Soybean/Ovule Stage/Nucellus
- Soybean Array/Soybean/test/Embryo Sac
- Soybean Array/SRB/Globular Stage/Embryo Proper
- Soybean Array/SRB/Globular Stage/Suspensor
Validation Experiments of Soybean GeneChip Data
Background
GeneChip has a limitation to detect mRNAs with low accumulation level. mRNA called "absent" by Affymetrix MAS5.0 can be detected by qRT-PCR, but at a reduced level. We determined that qRT-PCR is at least 1,000 times more sensitive than GeneChip (unpublished data). To identify mRNAs specific to different seed compartment during the development for RNAi study, we perform validation experiments for all compartment-specific transcription factors as well as for selected mRNAs with the highest GeneChip signal intensities. All compartment-specific candidates are detected in only one tissue and not detected in the other tissues at a specific soybean seed development stage.
Methods
1. Sample Preparation
- Using the same source of total RNA used for the GeneChip hybridization
- Using two biological replicates used for GeneChip analysis
- 2ng of total RNA from each seed compartment used for amplifying RNA
- Synthesizing single strand cDNA after one round of RNA amplification
2. Primer and PCR target design
- Same target sequences used by Affymetrix to design the probe sets on Affymetrix Soybean array.
- Using Beacon Designer software to design primers
- Primer length: 20~30-mer
- Tm of the primer: 60±5°C (mostly around 60±2°C, ΔTm<=5°C)
- The size of qRT-PCR product: 80-250bp (mostly 100-200bp)
3. qRT-PCR reactions
- Estimated 50pg of synthesized cDNA template used in each reaction
- (Two technical replicates X two biological replicates) = four reactions for each primer pair
- 18S rRNA as internal control for Ct normalization for each template
- 50 cycles amplification
- Melt curve analysis to evaluate Tm of qRT-PCR product
- Agarose gel analysis to evaluate the size of qRT-PCR product
Results
qRT-PCR experiments have validated the majority of compartment-specific transcription factor mRNA GeneChip data at the globular, heart, cotyledon, and early maturation stages of soybean seed development. In the following table, we carried validation experiments for 42 transcript mRNAs identified in soybean globular stage compartments (click the following link to download the pdf file).
Soybean Analysis Tools

- Click to browse the mRNA profiles of compartments during soybean seed developments by probe identification, gene ontology, or function category.

- Click to compare gene activity in different soybean seed compartments at different developmental stages.

- Click to BLAST DNA sequence against target sequences on the Affymetrix soybean array and view the seed expression pattern related to your gene-of-interest.
Soybean
Harvesting Seeds
Soybean plants (Glycine max cv. Asgrow A3237) were grown in the UCLA Plant Growth Center with a 16:8 light-dark cycle at 22°C. A linear correlation was observed between early developmental stages (globular to heart stage) and the length of soybean pods. Independent pod replicates at appropriate developmental stages were collected during the morning, and the seeds were staged by viewing dissected whole mount embryos using light microscopy. To increase precision in our measurements the pedicels were not included. Pods were placed immediately on ice and carried to the lab after collection.
Seed Fixation and Embedding
Seeds were dissected from pods no later than three hours after collection. A small excision was made on the chalazal end to allow fixative to penetrate seed tissues. After dissection, seeds were placed in a chilled solution of Acetic acid-Ethanol (3:1, v/v). Fixation was carried out for ~16 hours at 4°C. Seeds were dehydrated with a series of ethanol solutions with increasing concentration (75%, 85%, 95%, 3x 100%) for a minimum of 3 hours each. Seeds were then infiltrated with different xylene dilutions (in ethanol) (25%, 50%, 75%, 3x100%) for a similar period of time. Finally, seeds were incubated approximately 9-10 times with molten paraffin solution at 60°C for a minimum of 3 hours each change, embedded in paraffin in foil boats, and kept at 4°C until further use.
Seed Sectioning
Soybean seed paraffin sections of 6 µm and 8 µm were generated using a Reichert-Jung 820-II microtome. The paraffin ribbons were visualized using a compound microscope with a 10x magnification, and only ribbons containing visible embryos were placed on PEN-Foil slides. The slides were kept on a slide warmer at 45°C for approximately 24 hours, and then kept in a slide holder at room temperature to minimize exposure to humidity. A range of 5 to 8 slides per replicate was generated.
LCM
The PEN-Foil slides containing the paraffin sections were treated with a xylene solution in order to melt the paraffin. Two incubations of two minutes each were performed in the fume hood, where the slides were dried in the hood until the following day. The slides were then placed back in the slide holder and stored as previously mentioned in the Seed Sectioning section. The slides were used within two months of the xylene treatment.
We used either the Leica AS LMD or LMD6000 system to capture the different seed regions. Only sections containing a full embryo (i.e. embryo proper and suspensor) were captured and used for gene expression analysis. For each slide, the sections meeting this criterion were selected, a picture of the region to be captured was taken, the region was captured by LCM, and a final picture was taken documenting the captured area. Captured material was collected on a 0.5-ml Eppendorf tube cap containing 30 µl of the extraction buffer from the Arcturus PicoPure RNA isolation kit. After collecting all samples from a particular seed region on a slide, the tube containing the captured material was stored at room temperature until the end of the LCM session. Another tube was placed in the rack and a different seed region was captured from exactly the same sections that were used previously. At the end of the session, tubes containing the collected material were stored in a -80°C freezer until RNA isolation. As an example, one of the replicates with five slides contained 245 sections, but only 143 contained a full embryo from where the seed regions were captured. These 143 sections represented 36 seeds.
Also you can download the protocols below.