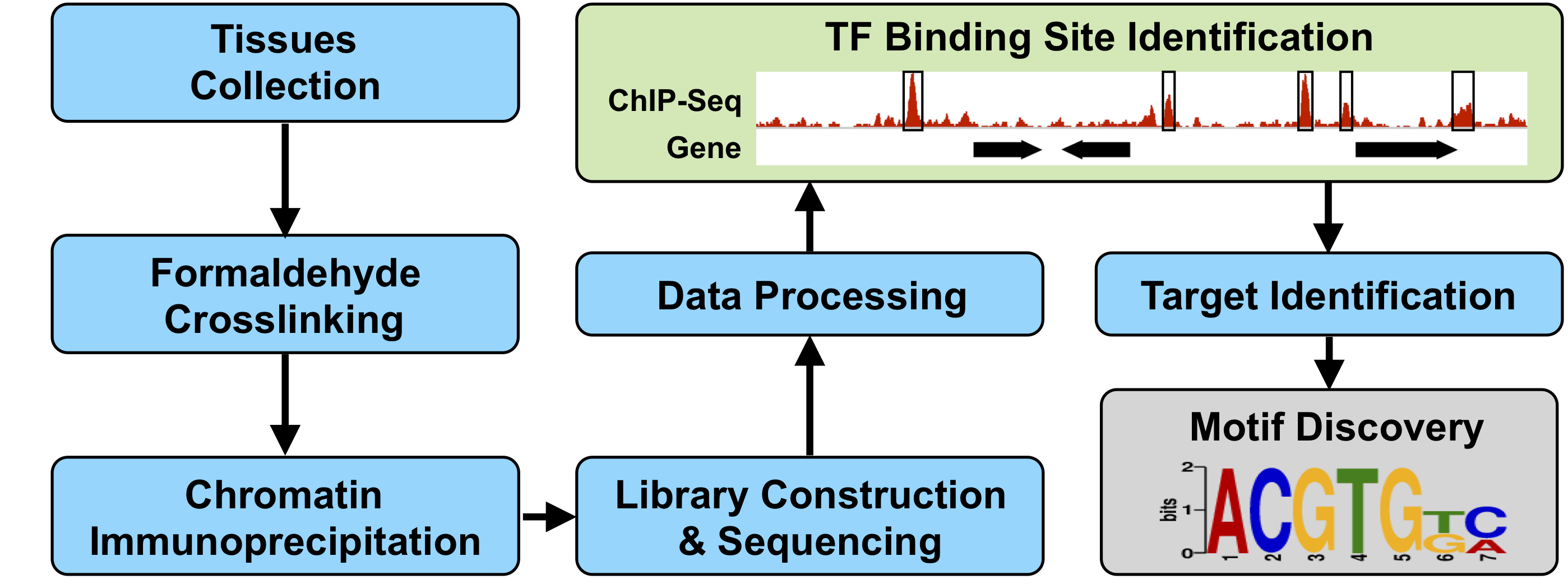
♦ Identifying Regulatory Networks That Control Seed Development Using DNA-Protein Interaction Profiling with Chromatin Immunoprecipitation (ChIP) Assays and Sequencing
We have generated a regulatory atlas of soybean seeds by identifying transcription factor (TF) mRNAs that are specific for every seed region and subregion throughout development (see transcriptome studies). A series of ChIP-Seq experiments are being carried out to determine (1) what downstream genes these TFs regulate, (2) what DNA regulatory motifs they interact with, and (3) how region- and subregion-specific TFs are organized into regulatory networks that program developmental events that give rise to a soybean seed.
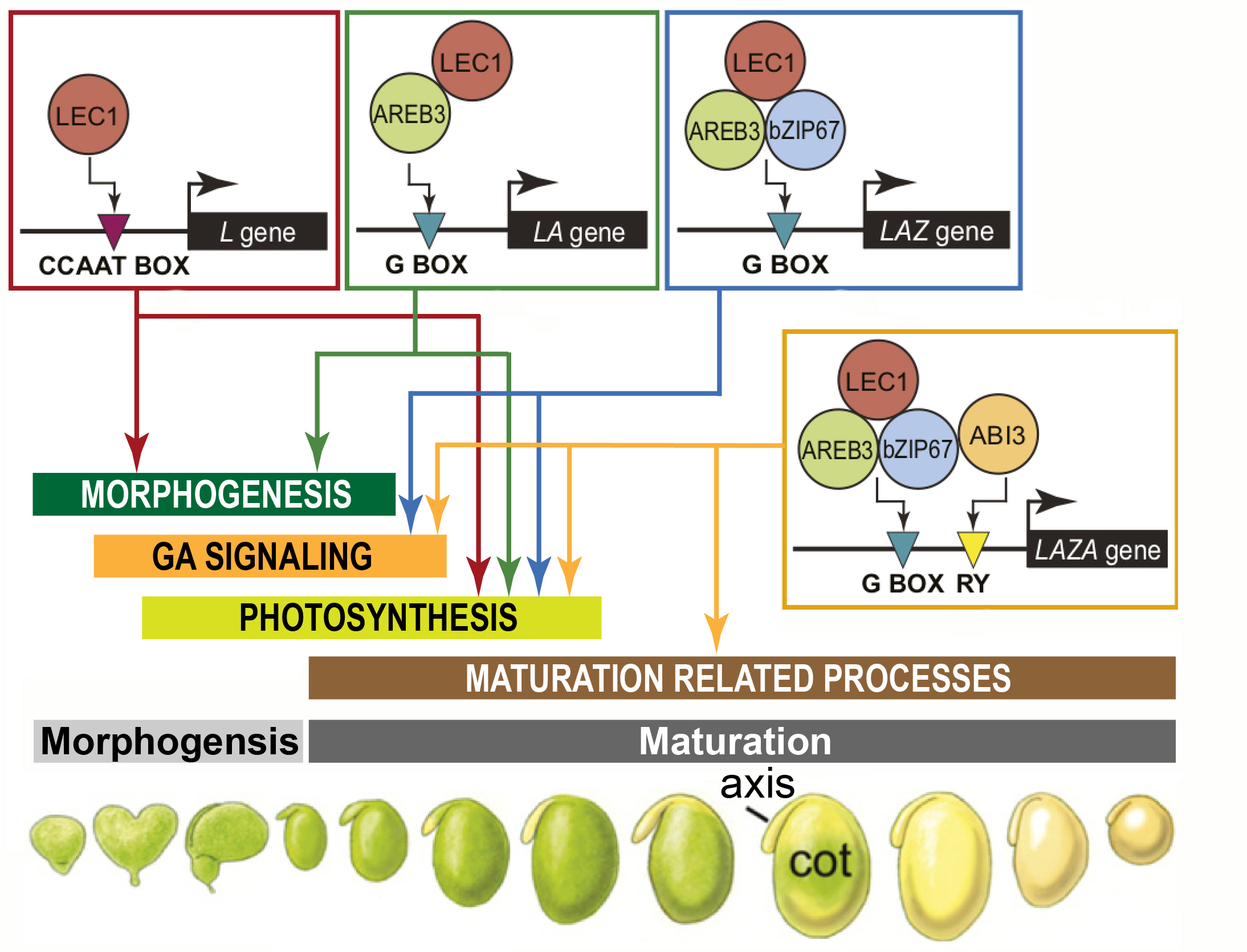
Experiments carried out at UC Davis with our collaborator, John Harada, investigated target genes that interact with one, or more, soybean TFs that program major developmental and physiological events during different periods of seed development. These include LEC1, AREB3, bZIP67, and ABI3 seed-specific TFs. These experiments showed that LEC1 regulates gene sets at different developmental stages and generates distinct biological processes by interacting with AREB3, bZIP67, and ABI3 TFs in specific combinations. DNA binding sites of target genes bound by these TFs are organized into cis-regulatory modules (CRMs) that are in close proximity to their target genes, and are enriched for DNA motifs known to bind to LEC1, AREB3, bZIP67, and ABI3 TFs. Early maturation stage embryo protoplast transcriptional assays validated that the CRMs are functional.
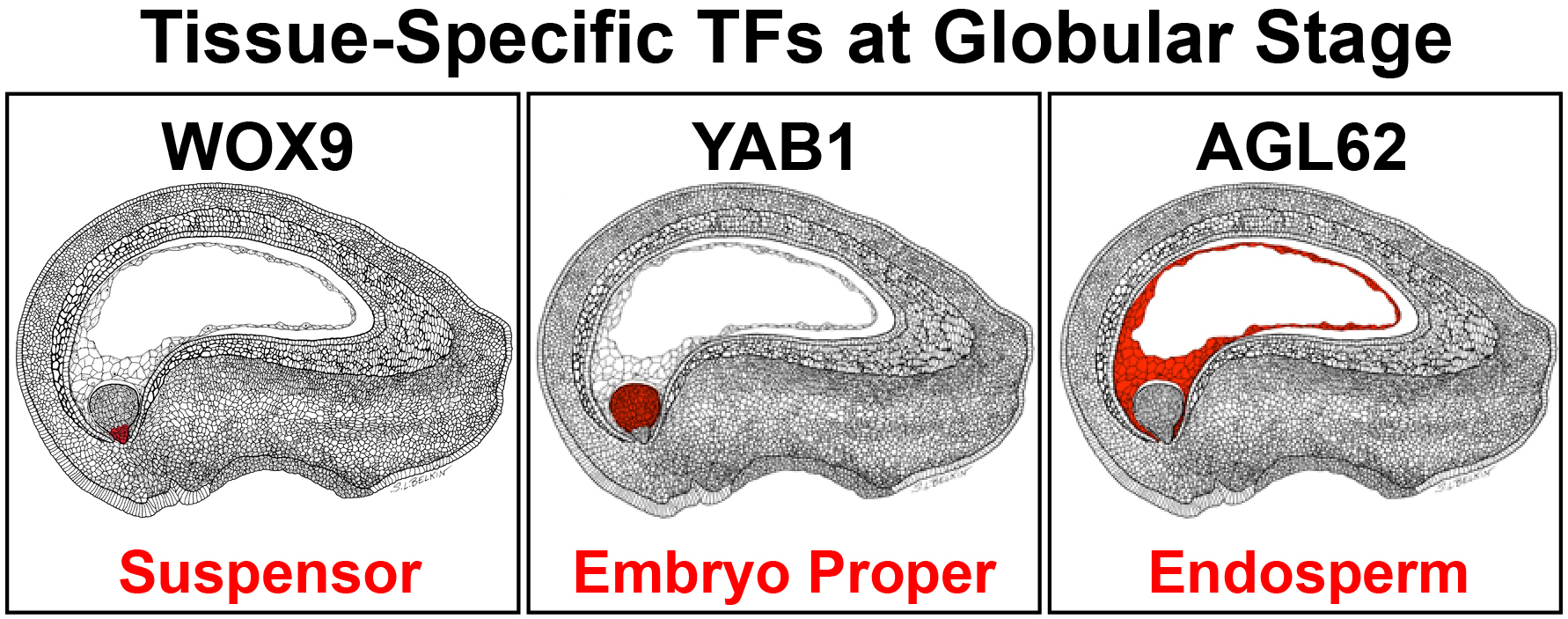
ChIP-Seq experiments carried out in our laboratory at UCLA identified target genes for TFs that are specific for different regions and subregions of soybean post-fertilization stage globular seeds. These include AGL62, YAB1, and WOX9 TFs that are specific for the endosperm, embryo proper, and suspensor, respectively. In addition, comparative genomic approaches are being used to identify embryo-proper- and suspensor-specific TFs present in seeds across the plant kingdom to uncover major regulatory pathways that program region-specific events during early seed development.
To date, we have generated 126 ChIP-Seq Datasets and submitted all to GEO. Click here to download the ChIP-Seq datasets.
Publications:
Jo, L., Pelletier, J.M., Hsu, S., Baden, R., Goldberg, R.B., Harada, J.J. (2020) Combinatorial interactions of the LEC1 transcription factor specify diverse developmental programs during soybean seed development Proc. Natl. Acad. Sci. USA, 117:2, 1223-1232.[DOWNLOAD PDF]
Pelletier, J.M., Kwong, R. Park, S., Le, B.H., Baden, R., Cagliari, A., Hashimoto, M., Munoz, M., Fischer, R.L., Goldberg, R.B., Harada J.J. (2017) LEC1 sequentially regulates the transcription of genes involved in diverse developmental processes during seed development. Proc. Natl. Acad. Sci. USA, 114:32, E6710–E6719. [DOWNLOAD PDF]
♦ Using Giant Bean Embryos To Dissect the Regulatory Networks That Control Early Embryo Development
The Scarlet Runner Bean (Phaseolus coccineus) provides a novel opportunity for dissecting the molecular processes controlling plant embryo development. At the globular stage, the Scarlet Runner Bean embryo is ~100-times larger than that of Arabidopsis, contains a suspensor with 200 cells that is highly polyploid, and can be isolated directly from developing seeds within the flower. Because of its large embryonic size, both embryo proper and suspensor regions can be separated from each other manually and used directly for biochemical and molecular studies. Almost 50 years ago, the late Ian Sussex and his collaborators pioneered the use of giant Scarlet Runner Bean embryos, and provided the first insights into the molecular processes controlling early embryogenesis; for example, the suspensor produces signals that are required for embryo proper development. During this same period, others demonstrated that hormones, such as gibberellic acid, are synthesized within the giant Scarlet Runner Bean suspensor and contribute to embryo proper formation.
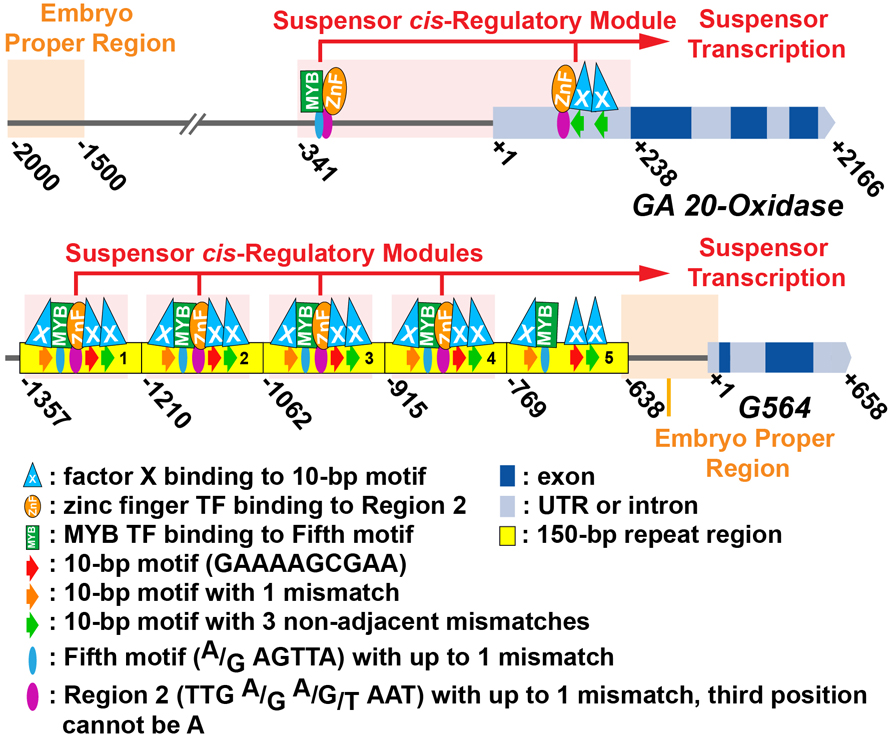
We analyzed the upstream region of two suspensor region-specific genes, G564 and GA 20-oxidase to understand how they are activated specifically within the suspensor during early embryo development. Fine-scale deletion, scanning mutagenesis, and gain-of-function experiments in transgenic tobacco embryos uncovered cis-regulatory elements required to activate transcription in the suspensor, including the 10-bp, Region 2, and Fifth motifs. Motif searches suggest that motifs in G564 and GA20-oxidase upstream regions interact with MYB and zinc finger TFs, respectively. Our results demonstrate the flexibility of motif positioning within a cis-regulatory module that activates gene transcription within giant bean suspensors and suggest that G564 and GA 20-oxidase comprise part of a suspensor gene regulatory network.
Publications:
Chen, M., Bui, A.Q., & Goldberg, R.B. (2020) Using giant scarlet runner bean (Phaseolus coccineus) embryos to dissect the early events in plant embryogenesis. in Plant embryogenesis: Methods and Protocols , M. Bayer, Ed. (Springer Science+Business Media, 2020), 2122, 205-222. [DOWNLOAD PDF]
Henry, K.F., Bui, A.Q., Kawashima, T., and Goldberg, R.B. (2018). A shared cis-regulatory module activates transcription in the suspensor of plant embryos. Proc. Natl. Acad, Sci. USA. (2018). 115 (25) E5824-5833. [DOWNLOAD PDF]
Henry, K.F., Kawashima, T., and Goldberg, R.B. (2015). A cis-regulatory module activating transcription in the suspensor contains five cis-regulatory elements. Plant Mol. Biol. 88, 207-217. [DOWNLOAD PDF]
Henry, K.F. and Goldberg, R.B. (2015). Using giant scarlet runner bean embryos to uncover regulatory networks controlling suspensor gene activity. Frontiers in Plant Science 6, 44, 1-6. [DOWNLOAD PDF]
Kawashima, T., Wang, X., Henry, K.F., Bi, Y., Weterings, K., and Goldberg, R.B. (2009). Identification of cis-regulatory sequences that activate transcription in the suspensor of plant embryos. Proc. Natl. Acad. Sci. USA, 106, 3627-3632. [DOWNLOAD PDF]
Le, B. H., Wagmaister, J.A., Kawashima, T., Bui, A. Q., Harada, J.J., and Goldberg, R.B. (2007). Using genomics to study legume development. Plant Physiology 144, 562-584 (doi.org/10.1104/pp.107.100362). [DOWNLOAD PDF]
Weterings, K., Apuya, N.R., Bi, Y., Fischer, R.L., Harada, J.J., and Goldberg, R.B. (2001). Regional localization of suspensor mRNAs during early embryo development. Plant Cell, 13, 2409-2425. [DOWNLOAD PDF]