Arabidopsis Seed Tissues/Compartments GeneChip Experiments.
We completed 65 GeneChip experiments of 31 tissues/compartments throughout Arabidopsis seed development using Affymetrix Arabidopsis ATH1 arrary (Click here to view our annotation of Arabidopsis ATH1 array).
Figure. Tissues/Compartments Studied Throughout Arabidopsis Seed Development (All images were illustrated by Meryl Hashimoto. The images are not to scale. Click on the image to enlarge.)
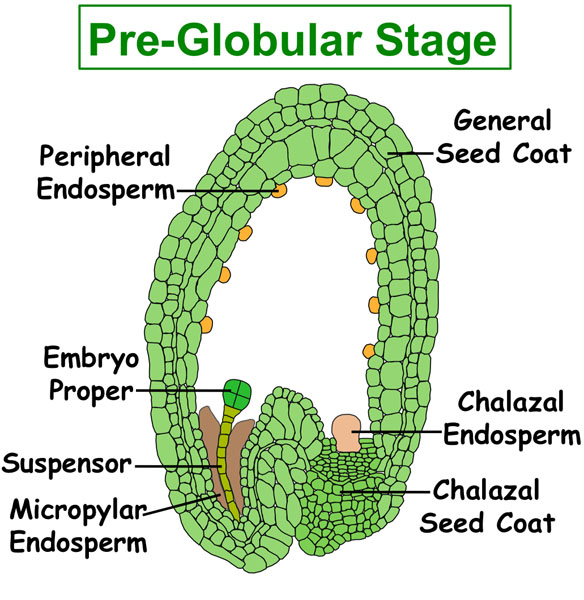 |
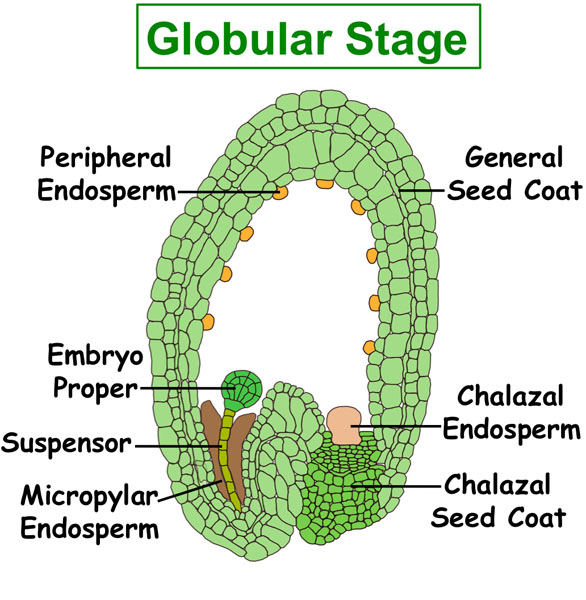 |
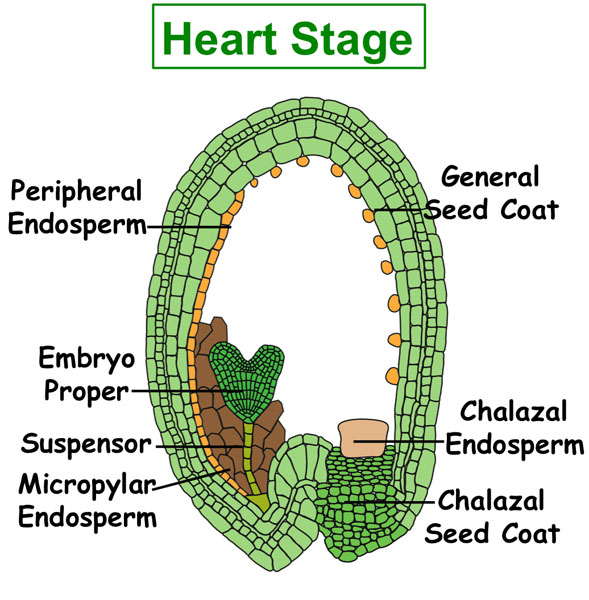 |
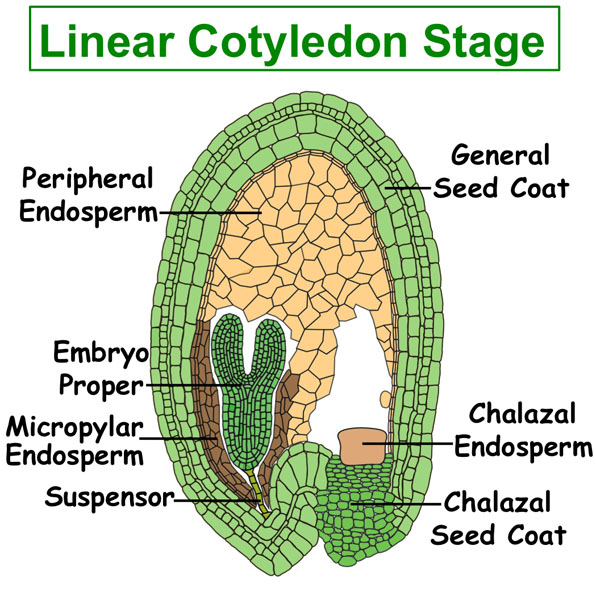 |
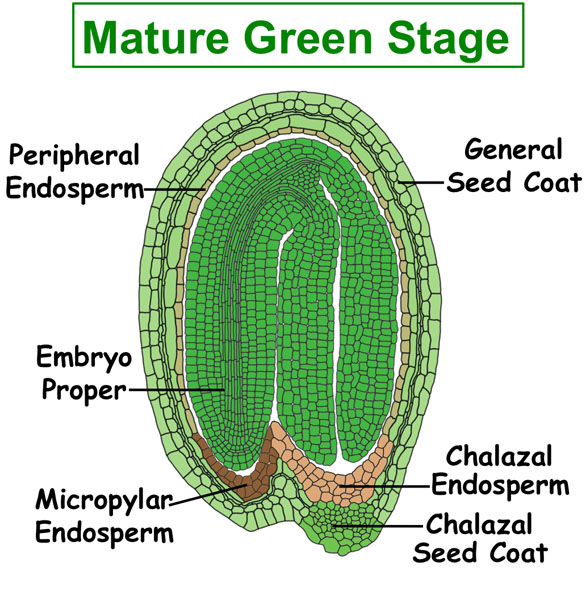 |
Click here to use our web-based analysis tools to analyze GeneChip experiments of arabidopsis seed tissues/compartments.
Click on the name of Arabidopsis seed tissues/compartment listed below to find the detail information about GeneChip experiments of Arabidopsis seed tissues/compartments. The information includes:
(1) GeneChip hybridization & analysis settings;
(2) Gene Expression Omnibus (GEO) accession;
(3) Images of sections captured;
(4) Growth conditions and protocols.
The notation for the experiment is as follows: GeneChip Array / Plant Species / Developmental Stage / Cell & Tissue Type.
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Chalazal Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Chalazal Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Embryo Proper
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/General Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Micropylar Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Peripheral Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Seed
- Arabidopsis ATH1 Array/Arabidopsis/Globular Stage/Suspensor
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Chalazal Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Chalazal Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Embryo Proper
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/General Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Micropylar Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Peripheral Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Heart Stage/Seed
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Cellularized Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Chalazal Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Chalazal Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Embryo Proper
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/General Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Micropylar Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Linear Cotyledon Stage/Seed
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Chalazal Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Chalazal Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Embryo Proper
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/General Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Micropylar Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Peripheral Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Mature Green Stage/Seed
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Chalazal Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Chalazal Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Embryo Proper
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/General Seed Coat
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Micropylar Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Peripheral Endosperm
- Arabidopsis ATH1 Array/Arabidopsis/Preglobular Stage/Seed
Validation Experiments of Arabidopsis GeneChip data
The validation experiments of Arabidopsis GeneChip data are in progress. Please check back later for more information. Thanks.
Arabidopsis Analysis Tools

- Click to browse the mRNA profiles of compartments during Arabidopsis seed development by probe identification, gene ontology, or function category.

- Click to compare gene activity in different Arabidopsis seed compartments at different developmental stages.

- Click to BLAST DNA sequence against target sequences on the Affymetrix Arabidopsis ATH1 array and view the seed expression pattern related to your gene-of-interest.
Arabidopsis
Harvesting Seeds
Arabidopsis Plants were grown in a Conviron chamber under continuous light with fluorescent lamps at 20°C and 50% - 70% relative humidity. Along the shoot, a linear correlation was observed between the developmental stages and the length of Arabidopsis siliques. Independent silique replicates at appropriate developmental stages were collected during the afternoon and the seeds were staged by viewing whole mount seeds using Normarski microscopy.
Seed Fixation and Embedding
Seeds were either dissected out of siliques or left in sub-divided siliques and fixed in 3:1 (v/v) ethanol to acetic acid. Fixation was performed for 4 hours at 4°C, washed 3 times with 70% ethanol for 5 minutes each and kept in 70% ethanol overnight. Seeds were dehydrated with a series of ethanol solutions with increasing concentration (85%, 95%, 100%) for a minimum of 1 hours each. Seeds were then infiltrated with different xylene dilutions (in ethanol)(25%, 50%, 75%, 100%) for a minimum of 2 hours each. Finally, seeds were incubated approximately 3-4 times with melted paraffin at 60°C for a minimum of 3 hours and embedded with paraffin in metal boats and kept at 4°C until further use. Each block contains 30-50 seeds.
Seed Sectioning
Arabidopsis seed paraffin sections of 5 µm and 7 µm were generated.The slides were kept in a oven at 42°C overnight.
LCM
The PEN-Foil slides containing the paraffin sections were treated with xylene solution in order to melt the paraffin. Two incubations of two minutes each were performed in the fume hood, where the slides were kept at room temperature until the following day. The slides were kept in a slide holder at room temperature to minimize exposure to humidity.
We used Leica LMD6000 system to capture the different seed regions. For each slide, the sections meeting the criterion for each compartment were selected, a picture of the region to be captured was taken, the region was captured by LCM, and a final picture was taken documenting the captured area. Captured material was collected on a 0.5 ml eppendorf tube cap containing 30 µl of RNAqueous lysis buffer from the Ambion RNAqueous-Micro kit. After collecting all samples from a particular seed region on a slide, the tube containing the captured material was immediately spun down and transferred to dry ice. At the end of the session, tubes containing the collected material were placed at -80°C until further use. As an example, we collected 895 sections for one replicate of Peripheral Endosperm, 376 sections for one replicate of General Seed Coat.
Also you can download the protocols below.
Arabidopsis
Figure 1. Longitude section of Arabidopsis seed development stages
Laser-capture microdissection of Arabidopsis seed compartments [Download video]