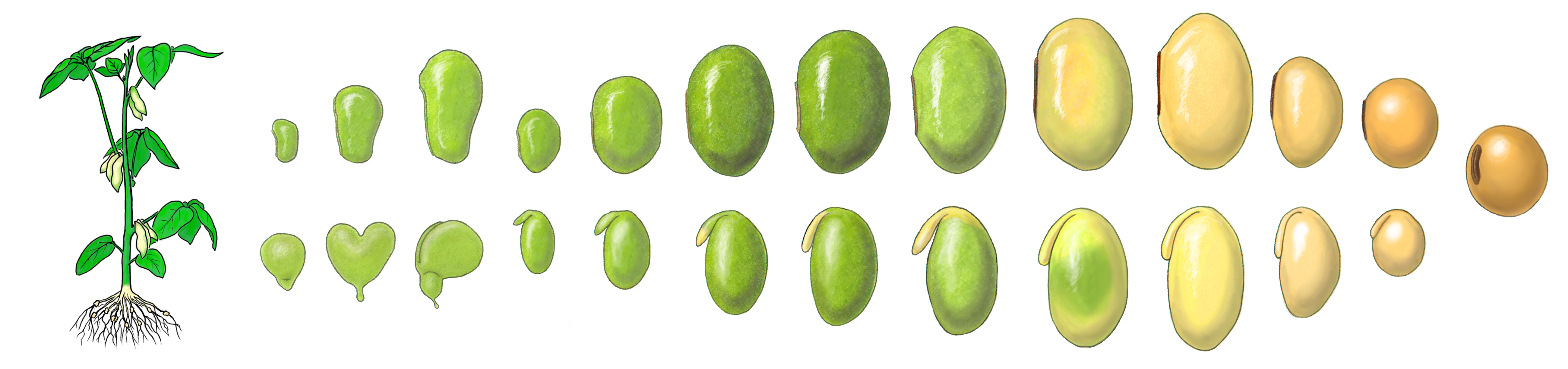
Welcome to Gene Networks in Seed Development Website!
Our long-term goal is to understand the genes and regulatory networks required to make a seed. Our National Science Foundation (NSF) Plant Genome Program funded research projects are a collaboration between the Goldberg and Pellegrini laboratories at UCLA, Harada laboratory at UC Davis.
Soybean is the most important oilseed crop in the world. Soybean seeds contain a high nutritional value, rich in proteins and oils, that serve as a major source of food for humans and animals. Soybean seed development is triggered by a double-fertilization process that leads to the differentiation of the embryo, endosperm, and seed coat that are the major regions of the seed and essential for seed viability and plant reproduction. Many different developmental and physiological events occur within each soybean seed region during development that are programmed, in part, by the activity of different genes.  Seed development, therefore, is the result of a mosaic of different gene expression programs occurring in parallel in different seed compartments. What these programs are and how they are organized into unique regulatory circuits within the plant genome to “make a seed“ remain major unanswered questions.
Seed development, therefore, is the result of a mosaic of different gene expression programs occurring in parallel in different seed compartments. What these programs are and how they are organized into unique regulatory circuits within the plant genome to “make a seed“ remain major unanswered questions.
All of the research and datasets presented in this website have been made possible by grants from the National Science Foundation's Plant Genome Program.
GENBANK DATASETS |
|

To date, we have generated ~600 datasets which have been submitted to NCBI GenBank, including: * Seed Transcriptome Studies: 246 RNA-Seq Datasets and 176 GeneChip Datasets. [DOWNLOAD DATASETS]* Seed Methylome Studies: 52 BS-Seq Datasets. [DOWNLOAD DATASETS] * Seed Regulatory Network Studies: 126 ChIP-Seq Datasets. [DOWNLOAD DATASETS] |
|
SEED TRANSCRIPTOME STUDIES |
|
|
♦ Profiling the Transcriptomes of Every Seed Region, Subregion, and Tissue Throughout Soybean and Arabidopsis Seed Development – An Atlas of Seed Gene Activity We used laser capture microdissection (LCM) to isolate specific soybean and Arabidopsis seed regions (e.g., seed coat, endosperm, and embryo) subregions (e.g., embryo proper and suspensor), and tissues (e.g., cotyledon adaxial and abaxial parenchyma) at different stages of development. RNA-Seq (soybean) and microarray (Arabidopsis) transcriptome profiling experiments were carried out to determine (1) the spectrum of genes that are active in different parts of the seed during development, (2) what transcription factors are localized in specific seed regions and subregions, and (3) the biological processes that are partitioned within a seed which may play important roles in seed differentiation and/or function. We profiled the mRNA sets present in 40 soybean and 42 Arabidopsis seed compartments captured using LCM - from shortly after fertilization through the early maturation stage of development. [READ MORE] |
|
SEED METHYLOME STUDIES |
|
|
♦ Studying the Role of DNA Methylation During Seed Development We profiled soybean and Arabidopsis methylomes from the globular stage through dormancy and germination. CHH methylation increases significantly during development throughout the entire seed, targets primarily transposable elements (TEs), is maintained during endoreduplication, and drops precipitously within the germinating seedling. By contrast, no significant global changes in CG- and CHG-context methylation occur during the same developmental period. An Arabidopsis ddcc mutant lacking CHH and CHG methylation does not affect seed development, germination, or major patterns of seed gene expression. These results suggest that CHH and CHG methylation does not play a significant role in seed development, or regulation of seed gene activity - including genes encoding major storage proteins. [READ MORE] |
|
SEED REGULATORY NETWORKS STUDIES |
|
|
♦ Identifying Regulatory Networks That Control Seed Development We have generated a regulatory atlas of soybean seeds by identifying transcription factor (TF) mRNAs that are specific for every seed region and subregion throughout development (see "Seed Transcriptome Studies" above). A series of ChIP-Seq experiments were being carried out to determine (1) what downstream genes these TFs regulate, (2) what DNA regulatory motifs they interact with, and (3) how region- and subregion-specific TFs are organized into regulatory networks that program developmental events which give rise to a soybean seed. [READ MORE] |
|
OUTREACH ACTIVITIES |
|
 Professor Goldberg created and currently teaches a novel course series that is a part of our NSF Plant Genome sponsored project. This outreach program contains a lecture course, HC70A - Genetic Engineering in Medicine, Agriculture, and Law and a lab course, HC70AL - Gene Discovery Lab. This program uses long-distance learning and targets non-science majors and entering life science students, in addition to students from underserved communities. Our objective is to teach undergraduates about the excitement of discovery, the process by which science is carried out, how advances in genetic engineering affect our daily lives, and how science is taught. The lecture part of this program (HC70A) is taught simultaneously between UCLA (Bob Goldberg), UC Davis (John Harada), and Tuskegee University (Channapatna Prakash). [Click here to learn more details about this unique teaching program.]
Professor Goldberg created and currently teaches a novel course series that is a part of our NSF Plant Genome sponsored project. This outreach program contains a lecture course, HC70A - Genetic Engineering in Medicine, Agriculture, and Law and a lab course, HC70AL - Gene Discovery Lab. This program uses long-distance learning and targets non-science majors and entering life science students, in addition to students from underserved communities. Our objective is to teach undergraduates about the excitement of discovery, the process by which science is carried out, how advances in genetic engineering affect our daily lives, and how science is taught. The lecture part of this program (HC70A) is taught simultaneously between UCLA (Bob Goldberg), UC Davis (John Harada), and Tuskegee University (Channapatna Prakash). [Click here to learn more details about this unique teaching program.]
|
|
COMMUNITY CONTRIBUTIONS |
|
 Our unique seed transcriptome and methylome datasets are also curated into SoyBase for the whole soybean community to use. The seed transcriptome datasets can be found under SoyBase Gene Expression Explorer page. The seed methylome datasets can be found under SoyBase Genome Methylation Explorer page.
Our unique seed transcriptome and methylome datasets are also curated into SoyBase for the whole soybean community to use. The seed transcriptome datasets can be found under SoyBase Gene Expression Explorer page. The seed methylome datasets can be found under SoyBase Genome Methylation Explorer page.
|
|
